The reticulate package allows R and Python to work together — here’s a tutorial
R and Python have many similarities and many differences. Most of the underlying concepts of data structures are very similar between the two languages, and there are many data science packages that now exist in both languages. But R is set up in a way that I would describe as ‘data first, application second’, whereas Python feels more application development driven from the outset. Javascript programmers, for example, would slot into Python a little quicker than they would slot into R, purely from a syntax and environment management point of view.
More and more I have been working in R and Python and I have come across situations where I’d like to use both together. This can happen for numerous reasons, but the most common one is that you are building something in R, and you need functionality that you or someone else has written previously in Python. Sure, you could rewrite it in R, but that’s not very DRY is it?
The reticulate package in R allows you to execute Python code inside an R session. It’s been around for a few years actually, and has been improving more and more, but it’s only recently that I’ve needed to use it, so I wanted to type up a brief tutorial on how it works. It you are an R native, getting reticulate up and running requires you to understand a little about how Python works — and how it typically does environment management — and so this tutorial may help you get it set up much quicker than if you tried to work it out yourself.
Environments in R and Python
Any programming project operates in an environment, which is where it stores and accesses all the things in needs or creates during its execution. In R, a common global environment is available to all projects, where the R base language and all installed packages are to be accessed. In this sense, all projects in R are usually run through the same common core environment. One way to think about this is to imagine that everyone in your house shares the same charging hub for their iPhones. They have to leave their room to charge the phone, and if they sell it the buyer will need to sort out their own charging arrangement.
In Python, however, each project is usually set up to be completely self contained — with its own environment, its own copy of the Python base and independent copies of all the modules it needs to execute. You can think about this as everyone having their own iPhone charger in their room. They don’t have to go outside and plug in somewhere else, and if they sold the phone, it comes complete with its own charger.
The Python model is more expensive in terms of installation processes and disk/memory resources, but it allows easier transfer of projects between individuals with minimal configuration, so it’s not hard to see how it has grown more directly out of a software development mindset, which is why I regard Python as more ‘application driven’.
Here’s a little graphic I sketched out to explain in simple terms the difference between how environments usually work in R and Python:
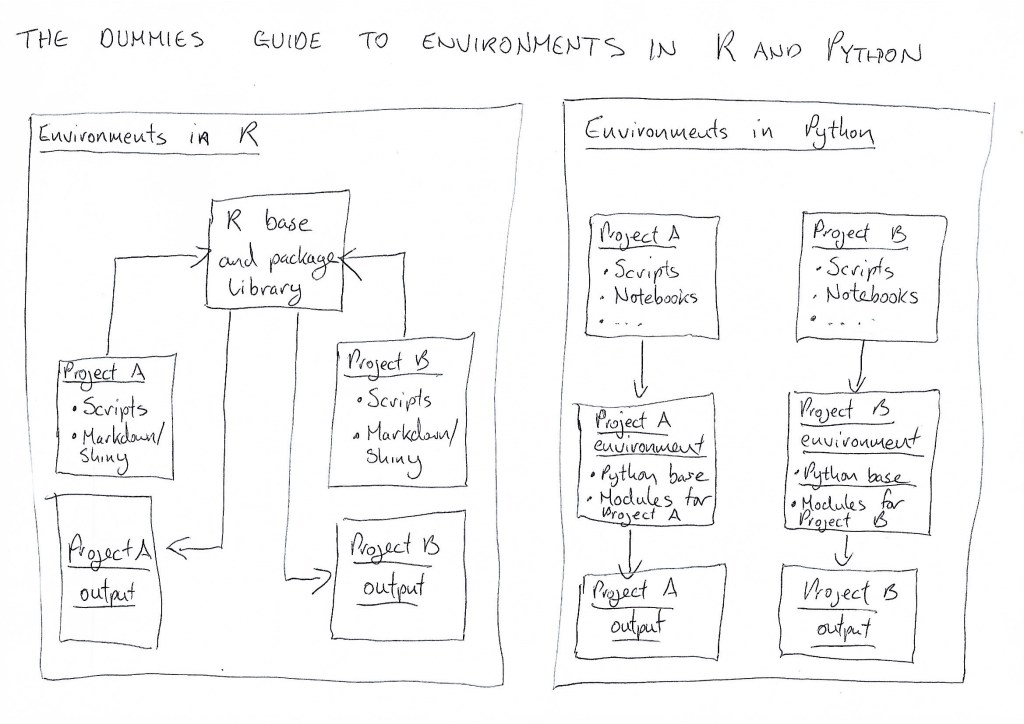
Now, if you want Python to talk to R, it still needs to find its environment — you can’t tell it to access R’s global environment. That would be like telling an English-speaker to find directions by asking a Chinese-speaker.
So, to get Python working inside your R project, you need two things:
- A Python environment set up inside your R project, so Python can get its bearings
- The
reticulatepackage to translate the Python code so that it works in R
Setting up a Python environment
From now on I am going to use a simple example. Let’s suppose I have an R project in RStudio which needs to use a function I have written in Python. So here’s a simple function which I will save in a Python script called light_years.py in my R project directory called test_python (yes, RStudio allows you to create Python scripts!). This function takes a distance in either kilometers or miles as an input and calculates how many years it would take to travel that distance at the speed of light — in other words, what is the distance in light years:
from scipy.constants import c
def light_years(dist, unit = "km"):
c_per_year = c * 60 * 60 * 24 * 365.25
if unit == "km":
dist_meters = dist * 1000
elif unit == "mi":
dist_meters = dist * 1.60934 * 1000
else:
sys.exit("Cannot use that unit!")
return dist_meters/c_per_yearI am using a very simple function example here to keep this article straightforward, so it’s a little unrealistic, and also a bit silly since I am importing the entire scipy package just to get the value of a constant, but hopefully it will help you get the idea.
Now as we discussed above, we need to provide this code with an environment. It needs:
- A version of Python to work through
- Access to the
scipypackage so it can get the constantc= speed of light
It’s not hard to set up a Python environment for your R project. Given how important project environments are in Python, numerous easy to use environment management tools exist.
My favourite is Anaconda. There are two versions available. The full version, contains a large universe of all the things an environment may need, including all the most used Python modules. Then there is Miniconda, which is easier on disk space and more appropriate for limited Python users. You can get Miniconda for your operating system here. Make sure you are downloading Conda for the version of Python that you want to work in.
Once you’ve installed Conda, if you are in MacOS or Linux, you’ll usually setup your environments using the command line. Just navigate to your R project directory (in my case test_python) in the terminal and use this command:
conda create --name test_pythonSimple as that, you now have a python environment created. I usually name my environments the same as the project folder to avoid future confusion.
Now you need to tell Conda to use that environment for this project, so while still in your test_python directory in the command line, use this command:
conda activate test_pythonAnd now you’ve linked this project to the Python environment, and there is a copy of the Python base in there for your code to run through.
Finally, our function needs the scipy package, so we will need to have that inside the environment. This is as simple as typing this inside the activated project folder:
conda install scipyConda will then install scipy and all dependencies it thinks it might need into your active environment and you are ready to go — easy as scipy, so to speak.
Now, later you are going to need to tell R where to find Python in this environment, so if you use this command, you can get a list of all environments and the path to where the environments were installed:
conda info --envsThis tells me, for example, that my environment was installed at /Users/keithmcnulty/opt/miniconda3/envs/test_python. I can always find the Python executables inside the bin subdirectory — so the full path to the Python executable for my project is /Users/keithmcnulty/opt/miniconda3/envs/test_python/bin/python3, since I am using Python 3. This is everything we need to tell R where to find the Python environment.
Running your Python function in R
Now, whether you’ve set up your Python environments like I did using Conda, or whether you have used virtualenv, you’ve done the hard bit . The rest is straightforward because reticulate takes care of it.
First, you need to tell R where to find the Python executable in the right environment when it loads your project. To do this, start up an empty text file and add the following, replacing my path to whatever path matches your Python executable inside the project environment you created.
Sys.setenv(RETICULATE_PYTHON =
"/Users/keithmcnulty/opt/miniconda3/envs/test_python/bin/python3")Now save this text file inside your project directory with the name .Renv. This is a hidden file that R will execute whenever you start up your project in RStudio. So now shut down RStudio and restart it with your test_python project open and it will now be pointing to the Python environment.
Now if you haven’t already installed the reticulate R package, you should do so at this point. Once installed, you can try a few tests in the terminal to see if everything is as it should be.
First you can test if R knows where Python is. reticulate::py_available() should return "TRUE". You can also test if the Python modules you need are installed: reticulate::py_module_available("scipy") should return "TRUE". Assuming all that works, you are ready to bring your function into R.
You can source your Python script with a simple:
reticulate::source_python("light_years.py")Now you have the light_years() function available as an R function. Let’s see how many years it would take to travel a quadrillion miles at the speed of light:
> light_years(1000000000000000, "mi")
[1] 170.1074Nice! Obviously this is a very simple example but it does tell you all you need about how to integrate Python code into your R script. You can now imagine how you can bring in all sorts of functionality or packages that are currently Python-only and get them working in R — very exciting. For example, I recently needed to use a new graph community detection algorithm called leidenalg for which an implementation only currently exists in Python, but all my existing project code was in R. So I was able to use reticulate just like I did here to solve this problem.
To learn more about using Anaconda or Miniconda to set up Python environments, the user guide is here. To learn more about the wide variety of functionality available to translate Python to R, there’s a goodreticulate vignette here.

